Protein Homologs
The JAK3 protein has homologs that have been identified in six species besides human; all six of these non-human protein homologs are also named JAK3. The following links (accession numbers) can be used to see more information on each protein. For information about DNA homologs please click HERE. [1]
Human (Homo sapiens): NP_000206
-1124 amino acids
Chimpanzee (Pan troglodytes): XP_512502
-1094 amino acids
Dog (Canis familiaris): XP_852473
-1103 amino acids
Mouse (Mus musculus): NP_034719
-1100 amino acids
Rat (Rattus norvegicus): NP_036987
-1100 amino acids
Chicken (Gallus gallus): NP_990327
-1106 amino acids
Zebrafish (Danio rerio): XP_002663133
-1103 amino acids
Human (Homo sapiens): NP_000206
-1124 amino acids
Chimpanzee (Pan troglodytes): XP_512502
-1094 amino acids
Dog (Canis familiaris): XP_852473
-1103 amino acids
Mouse (Mus musculus): NP_034719
-1100 amino acids
Rat (Rattus norvegicus): NP_036987
-1100 amino acids
Chicken (Gallus gallus): NP_990327
-1106 amino acids
Zebrafish (Danio rerio): XP_002663133
-1103 amino acids
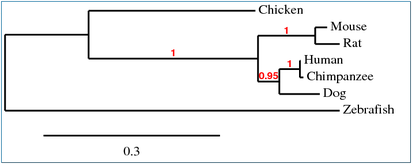
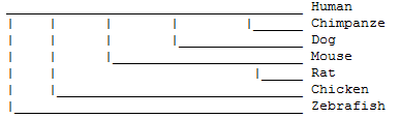
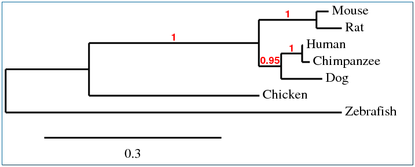
Phylogenetic Trees
The phylogenetic trees produces using MUSCLE, T-COFFEE and ProbCons (as represented in figure 1 and 2) represent the same information. The numbers on the figures indicate high confidence for the associated clade, where 1 means perfect confidence. The tree produced using TreeTop (figure 3) represents the same phylogenetic relationships between protein homologs, but the resulting figure is not as easy to read. The phylogeny website used to produce figures 1 and 2 are designed for easy use while still providing many options. The results are readable and can be downloaded in multiple formats, making it preferable over TreeTop for producing phylogenetic trees.
Overall these trees indicate that of the homologs, the chimpanzee JAK3 is the most similar to the human JAK3. The dog JAK3 is the next most similar to humans after the chimpanzee and the mouse and rat JAK3 are of equal similarity to human JAK3, but less similar than the dog. The chicken JAK3 is less similar to human JAK3 than the chimpanzee, dog, mouse and rat. The zebrafish JAK3 is the least similar of the homologs.
Alignments
| T-COFFEE protein alignment |
| GeneBee protein alignment |
| CLUSTALW protein alignment |
The alignments, as compared with the phylogenetic trees are useful because they allow users to see amino acid by amino acid where and how homologs differ.
All three protein alignments are useful, but the color coding used in the T-COFFEE [4] alignment makes it particularly easy to use compared with the GeneBee [3] and CLUSTALW [5] formats. The color coding makes it easy to locate areas of high or low homology across all seven homologs.
All three protein alignments are useful, but the color coding used in the T-COFFEE [4] alignment makes it particularly easy to use compared with the GeneBee [3] and CLUSTALW [5] formats. The color coding makes it easy to locate areas of high or low homology across all seven homologs.