MOTIF search
Using MOTIF Search on both the gene sequence and the mRNA sequence (as taken from NCBI, accession numbers available here), yielded different results. The mRNA alone yielded nine motifs while the gene yielded thirteen, indicating there are four motifs located in the non-coding portions of the gene.
mRNA found motifs
EGF_1: EGF-like domain signature 1
Prosite: PS00022
INTEGRIN_BETA: Integrins beta chain cysteine-rich domain signature
Prosite: PS00243
CTCK_1: C-terminal cysteine knot signature
Prosite: PS01185
ANAPHYLATOXIN_1: Anaphylatoxin domain signature
Prosite: PS01177
4FE4S_FERREDOXIN: 4Fe-4S ferredoxins, iron-sulfur binding region signature
Prosite: PS00198
THOILASE_3: Thiolases active site
Prosite: PS00099
TUBULIN: Tubulin subunits alpha, beta, and gamma signature
Prosite: PS00227
VWFC_1: VWFC domain signature
Prosite: PS01208
2FE2S_FER_1: 2Fe-2S ferredoxins, iron-sulfur binding region signature
Prosite: PS00197
Prosite: PS00022
INTEGRIN_BETA: Integrins beta chain cysteine-rich domain signature
Prosite: PS00243
CTCK_1: C-terminal cysteine knot signature
Prosite: PS01185
ANAPHYLATOXIN_1: Anaphylatoxin domain signature
Prosite: PS01177
4FE4S_FERREDOXIN: 4Fe-4S ferredoxins, iron-sulfur binding region signature
Prosite: PS00198
THOILASE_3: Thiolases active site
Prosite: PS00099
TUBULIN: Tubulin subunits alpha, beta, and gamma signature
Prosite: PS00227
VWFC_1: VWFC domain signature
Prosite: PS01208
2FE2S_FER_1: 2Fe-2S ferredoxins, iron-sulfur binding region signature
Prosite: PS00197
Gene found motifs*
IGF_BINDING: Insulin-like growth factor binding proteins signature
Prosite: PS00222
I_CONOTOXIN: I-superfamily conotoxin signature
Prosite: PS60019
AGOUTI_1: Agouti domain signature
Prosite: PS60024
DEFENSIN: Mammalian defensin signature
Prosite: PS00269
Prosite: PS00222
I_CONOTOXIN: I-superfamily conotoxin signature
Prosite: PS60019
AGOUTI_1: Agouti domain signature
Prosite: PS60024
DEFENSIN: Mammalian defensin signature
Prosite: PS00269
* only those motifs which were found in addition to those found using the mRNA sequence are listed.
Prosite motif search
MOTIF search provided motif information linking out to Prosite, but interestingly, when using Prosite to do the motif search on the JAK3 mRNA returned different results. The Prosite search with mRNA returned 11 motifs, including the 9 identified in the MOTIF search of the mRNA. The Prosite search, however, also returned the I-CONOTOXIN and DEFENSIN motifs identified in the MOTIF search using the gene sequence.
A Prosite motif search using the gene sequence returned thirteen motifs. The motif that the MOTIF search identified as IGF_BINDING (PS00222) was also located by Prosite but was idetified as IGFBP_N_1 (PS00222).
A Prosite motif search using the gene sequence returned thirteen motifs. The motif that the MOTIF search identified as IGF_BINDING (PS00222) was also located by Prosite but was idetified as IGFBP_N_1 (PS00222).
MEME motif search
| MEME discovered motifs |
Using MEME to search for motifs in the mRNA sequence found twenty motifs, all of which were present in the gene in at least two places. MEME is helpful for locating potential motifs but does not appear to be as helpful as MOTIF Search in identifying them. See the file at left for a screenshot of some of the MEME motif search results.
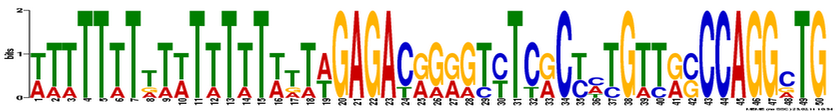
Align Ace motif search
A motif search using Align ACE was also conducted, and located 25 motifs. The Align ACE was probably the least helpful of the motif search databases as the output is limited to a list of the found motif sequences and a corresponding map score. See the file to the right for the Align ACE search results.
| Align ACE JAK3 motif search results |